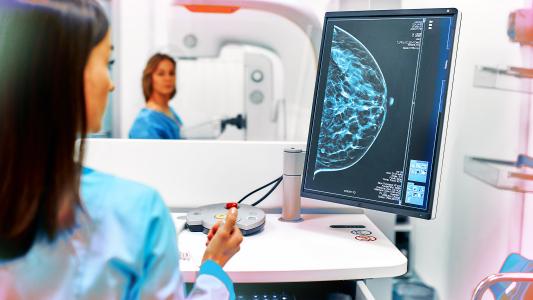
Cancer is measured in stages and grades. The stage describes the size of the tumor and how far it has spread. Meanwhile, grade describes the appearance of the cancerous cells compared to normal cells.
The stage and grade help doctors determine the best course of treatment and predict the risks for the patient, while doctors are increasingly turning to genetic sequencing to look for unique mutations that may be driving the cancer. However, recent research into epigenetics — broadly speaking, changes to gene expression that occur without changes to the DNA sequence itself — suggests epigenetic factors also affect one’s susceptibility.
“Traditionally, cancer has been viewed as primarily a result of genetic mutations,” Hilary Coller, a professor of molecular, cell, and developmental biology at UCLA, said. “However, the emergence of advanced next-generation sequencing technologies has made more people realize that the state of the chromatin and the levels of epigenetic factors that maintain this state are important for cancer and its progression.”
Coller and a group of researchers from the UCLA Health Jonsson Comprehensive Cancer Center wanted to determine how these factors affect cancer progression. Their findings suggest that epigenetic factors may not only predict patient outcomes better than traditional measures, they also lay the groundwork for artificial intelligence that will help doctors do so across multiple types of cancer.
Determining the epigenetic prospects
The researchers started by getting RNA sequencing data from various kinds of cancer in publicly available databases, such as The Cancer Genome Atlas (TCGA). They analyzed the data on the 24 adult cancer types to look for patterns among 720 epigenetic factors. Specifically, they looked at factors that encode proteins involved in the “addition, removal, and recognition” of DNA. They then clustered the cancers based on distinct expression patterns.
The researchers found that 10 of the 24 cancers they looked at had significant differences in at least one of three clinical outcomes: progression-free survival, disease-specific survival, and overall survival. In other words, looking only at the epigenetic factors, the researchers could predict if the patient ultimately had a better outcome in one of those three metrics.
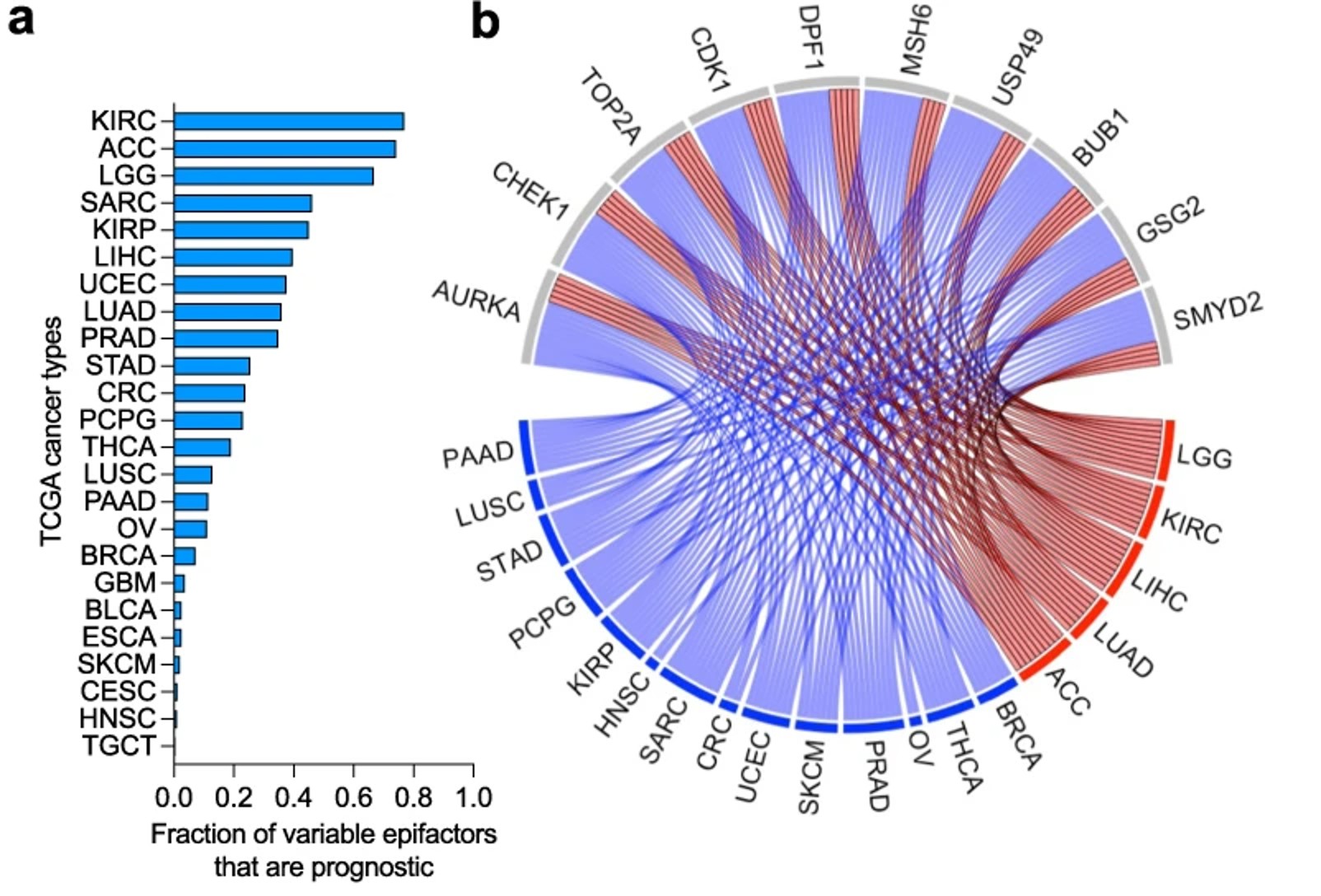
Additionally, five of the 10 showed significant predictive power in all three outcomes. Those cancers were from areas of the body as diverse as the liver, lungs, brain, and kidneys. The fifth was a cancer that originates in the adrenal glands (located above the kidneys).
Considering that grade-based forecasts are typically only effective at predicting outcomes for one or two cancer types out of the five — and across all three metrics — this suggests that epigenetic factors could help improve more traditional measurements.
“We saw that the prognostic efficacy of an epigenetic factor was dependent on the tissue of origin of the cancer type,” Mithun Mitra, an associate project scientist at the Coller laboratory and another co-senior author, said. “We even saw this link in the few pediatric cancer types we analyzed. This may help decide the cancer-specific relevance of therapeutically targeting these factors.”
AI predictions
The researchers next wanted to see if these epigenetic relationships could be sussed out by an AI. They used Cox-nnet, an artificial neural network specifically designed to predict patient outlooks, and focused on the five cancer types found to have the strongest predictive power. They trained the AI on 80% of the patient data and tested the resulting model on the remaining 20%.
“The [AI model] would be good to test it on other independent datasets to explore its broad applicability,” Mirta said. “Similar epigenetic factor-based models could be generated for pediatric cancers to see what factors influence the decision-making process compared to the models built on adult cancers.”
The results were successful. The AI was able to predict the clinical outcomes of the test cohort. The researchers further validated their model by testing it on other datasets. These tests also showed that the AI could use epigenetic factors to accurately determine which patients had a higher chance of good outcomes and which had a higher chance of poor outcomes.
The researchers published their findings in Communications Biology, a peer-reviewed, open-access journal.
The roadmap demonstrates how to identify certain influential factors in different types of cancer.
Michael Cheng
A roadmap for survival
In the study, the researchers suggest that a subset of specific epigenetic factors may work together to modulate tumor-suppressing genes. Being able to recognize those factors can help doctors predict patient outcomes and also present the starting line for developing targeted therapies. The speed at which AI may be able to analyze public datasets for epigenetic patterns can be a further benefit.
However, this study is preliminary. As the researchers note, their list of 720 epigenetic factors — while large — is hardly exhaustive and doesn’t take into account some known epigenetic mechanisms. Additional research into pediatric cancers specifically will be necessary to enhance medical understanding of their epigenetic factors, too.
Even so, as Michael Cheng, a graduate student in the Bioinformatics Interdepartmental Program at UCLA and the study’s first author, points out: The study offers a roadmap for future research as well as the improvement of AI models that can better guide treatment and, hopefully, boost survival.
“The roadmap demonstrates how to identify certain influential factors in different types of cancer and contains exciting potential for predicting specific targets for cancer treatment,” Cheng said.